Image

Building a mathematical model to artificially recreate proteins? The work of Rémi Monasson and Simona Cocco, researchers at the ENS Physics Laboratory (LPENS), shows how protein models, derived from natural amino acid sequence data, can be used to artificially design functional proteins, whose properties are tested and validated in vivo. A promising discovery for evolutionary biology and biomedical applications, published by the prestigious scientific journal Science.
Protein modeling: a crucial issue in evolutionary biology and biomedical applications
The design of proteins is of fundamental interest in biology but also in medicine or pharmacology. These molecules are entirely determined by the chain - the sequence - of amino acids that compose them. There are only twenty different types of amino acids in nature, but these alone code for the tens of thousands of proteins that keep us alive. These proteins differ from each other only in their number and sequence of amino acids.
Each protein acquires a particular shape in space under the effect of the forces between its amino acids. This is called the "folding" phenomenon. This shape gives it the ability to perform a specific task in the body, such as participating in the transport of oxygen, the immune response, the digestion process or the construction of muscles.
How then can we understand or predict the shape of a protein if we know its sequence? If the problem can be formulated in one line, its resolution is extremely complex and largely unfinished, although the applications are incredibly numerous.
As Rémi Monasson, a researcher in the Statistical Physics and Inference for Biology team at LPENS, explains, in the medical field or in pharmacology "it is crucial to understand how proteins are affected by amino acid changes occurring in genetic diseases, or how to design new ones with desired properties for therapeutic purposes."
An issue at the heart of the work of the scientist and his team, who thanks to applied methods of physics, managed to model synthetic proteins from amino acid sequences of various organisms. "The approach we followed to create these proteins is inspired by nature, and more precisely by the diversity of solutions it offers. Over the course of evolution, nature has generated numerous protein sequences," explains Simona Cocco, a researcher in the same team and one of the co-authors of the article published in Science.
Image
Combining the cutting-edge principles of statistical physics with the incredible resources of nature
For their work, the two scientists used a particular protein composed of about 100 amino acids: Chorismate Mutase (CM). Present in bacteria, fungi and plants - but not in animals, which can only obtain it through their diet - it is itself essential for the production of certain amino acids.
These organisms are so far apart in the evolutionary tree that the amino acid sequences of the Chorismata Mutases they contain in their DNA are very different from each other, even though they "code" for proteins with the same role.
The researchers used the thousands of sequences of this protein available in the databases to establish a mathematical model. This model assigns to each possible amino acid sequence a score, i.e. a probability, that it codes for a "good" protein, possessing the same properties as the natural CMs.
This model is inspired by the applied principles of statistical physics, where we assume that amino acids interact two by two, like elementary physical objects (electrons, spins, ...). The values of these interactions are not known a priori, but are calculated by requiring that the natural sequences have high scores.
Once this mathematical model was established, the team of researchers used it to create, on computer, new amino acid sequences with high scores. "These artificial sequences turned out to be very different from the natural sequences from which we learned our model. However, the proteins defined in this way are perfectly valid and have the same function as the natural CM proteins," says Rémi Monasson enthusiastically.
To test the compatibility of this new CM protein, the researchers used the bacterium E. coli, which is widely studied in biology. They first deleted the part of the DNA encoding the CM protein, which they then replaced with a portion of DNA corresponding to the computer-generated amino acid sequence. The result? These genetically modified bacteria behave like their natural counterparts, they grow and reproduce without difficulty.
A scientific victory, which in time should lead to important advances in medicine, both in the understanding of certain generic diseases and in their treatment.
"Our study provides evidence of the potential of protein modeling based on sequences from various organisms. It also shows that beyond the protein sequences that emerged by chance during evolution, a very large number of other sequences, which we have estimated in our work, are equally good solutions in terms of proteins," says Simona Cocco.
Thus, characterizing upstream this space of "good" sequences should allow a better understanding of how evolution has progressively explored it over billions of years. But for the researchers, much remains to be done: "among other things, it would be very important to be able to separate the different contributions to the functioning of a protein in our models, which would make it possible to act on a precise component, for example biochemical activity, without modifying the others, such as specificity or thermodynamic stability. In other words, the ultimate goal is to "break" the molecular code linking the amino acid sequence to the function of the proteins, which will make it possible to manipulate and modify the proteins at will.
Image

Interdisciplinarity, a vital approach to conceptual and technical progress
Our subject has been of great interest for more than fifty years to biologists, but also to chemists, computer scientists and physicists such as Simona Cocco and Rémi Monasson: "Proteins are extraordinary objects, at the junction between physics, chemistry and biology. They are clearly not alive and therefore belong to the sciences of matter. But the fact that they are capable of evolving, in the Darwinian sense of the term, unlike standard physical objects such as electrons, atoms, molecules, etc., makes them true biological objects at the frontier of physics.
Moreover, the complexity of modeling proteins and the limited success of usual approaches (such as writing energy as a function of elementary parameters, which are difficult to choose and estimate) makes their study particularly interesting for researchers.
According to Rémi Monasson, "we talk a lot about complex systems in physics, without this concept being always clearly defined. Proteins are definitely part of it... It is therefore necessary to invent new ways to study them. We can hope in return that the conceptual and technical progress they will generate will benefit other systems in physics. Interdisciplinarity in the study of proteins but also more broadly in biology is necessary for the researcher, "especially to meet the need for tools and digital methods to model, analyze and organize experimental data."
From his point of view, the contributions of computer science, mathematics and theoretical physics within the life sciences will be crucial in the decades to come: "I think that the main effort must be made in the training of students, who will obviously be the researchers of tomorrow. We must offer them the possibility of learning at the highest level in each of these disciplines and avoid at all costs conceiving interdisciplinarity as a superficial veneer. These training courses must enable these researchers to be recognized as physicists by physicists and biologists by biologists, and not the other way around... There is clearly a great deal at stake in this area in the years to come.
For Simona Cocco and Rémi Monasson, this goes hand in hand with the possibility of working in an environment that is conducive to exchanges and collaborations: "We very much appreciate the exceptional position of the ENS in the heart of Paris, surrounded by all the research institutions in the Île-de-France region, with which we can interact easily. We also appreciate the excellence of the students we work with in teaching, but also in internships and theses, which contributes to the quality and diversity of the research done here. In our field, statistical physics, it is difficult to think of any other place in the world with so many players and activities," conclude the two researchers.
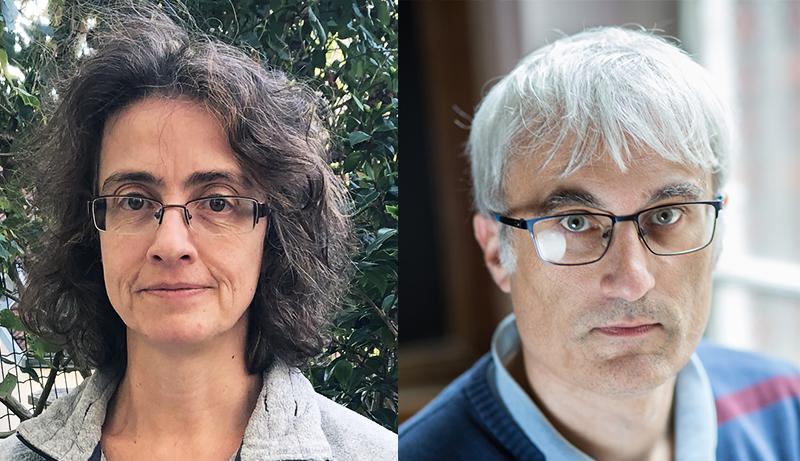
About SimonaCocco
Simona Cocco has been a research director at the CNRS since 2013 and a member of Section 5 of the National Committee for Scientific Research since 2018. After studying at the University of Rome La Sapienza, the Franco-Italian researcher obtained a double doctorate in physics and biophysics, following a thesis at the interface between physics and biology in cotutelle between the Department of Biophysics at La Sapienza and the Department of Physics at the École Normale Supérieure in Lyon.
She then did a post-doc at the UIC in Chicago, before becoming a CNRS research fellow at the Complex Fluid Laboratory in Strasbourg. In 2004, she joined the ENS-PSL physics department. From 2009 to 2011, she was a Senior Member at the Institute for Advanced Study at Princeton University.
"I first became interested in applications of statistical physics to neural networks, thanks to the courses of D. Amit, G. Parisi, M. Virasoro at La Sapienza. I then worked on the modeling of DNA micro-mechanics and on single molecule experiments in collaboration with J. Marko, and V. Croquette. Then, I focused my research on model inference from biophysical, genetic and neurosciences data. The increasing amount of data in biology as well as in other domains requires more and more analysis tools crossing statistical physics, inference and computational science.
About RémiMonasson
Rémi Monasson (Sciences 1988), a former student of the Ecole Normale Supérieure, quickly switched from mathematics to physics. He did his PhD at the ENS, in statistical physics under the supervision of Marc Mézard in (1993), where he started working on interdisciplinary applications of statistical physics, notably on neural networks and combinatorial optimization problems.
He continued as a postdoctoral fellow at the University of Rome La Sapienza for two years, and upon his return became a research fellow at the CNRS in the theoretical physics laboratory of the ENS. He then left for a stay at the UIC in Chicago and then in Strasbourg for 3 years. Since 2004, he has been working in the physics department of the ENS, except for a two-year stay in Princeton from 2009 to 2011, where he turned to systems biology following his meeting with S. Leibler. Rémi Monasson is also a professor of physics at École Polytechnique and deputy director of the Institut Henri Poincaré since 2018.
"I have always been attracted by the universal character of statistical physics, in the sense that its problematic - how to understand the emergent properties of a large number of elementary components? - goes far beyond the framework for which it was invented - understanding the properties of gases and thermodynamics - and arises in all sciences. Protein modeling is, along with neuroscience (specifically, how is space represented in the brain?) one of the two big questions I am interested in in biology."

Be the first to review this item, please login or register.
Sign-inRegister